Installing fcm flow cytrometry library on Windows 8.1
2014-05-14 16:28
My friend Andrew is working with flow cytrometry, which is a fancy way of saying “a field I don’t understand.” He wanted to use a python library for analyzing data, but was having trouble getting it installed. After some effort, I found this worked. The core problem is that py-fcm hasn’t been updated to work with later versions of its dependencies. It also doesn’t list the versions of its dependent libraries, making installation difficult.
Fortunately, I happened across a site that maintains Windows binaries for lots of packages, plus a science-ready Python distribution. I don’t know if this will install properly on Windows 7, but believe it will.
Downloads
Enthought Canopy Express
This is a free Python distribution. I installed the x64 version, specifically 1.4.0-win-64, which used Python 2.7. It includes the following packages that py-fcm depends on: numpy, scipy, matplotlib, cython.
https://www.enthought.com/downloads/
DPMix
The Windows installer version of DPMix. I used version dpmix-01.win-amd64-py2.7.exe.
http://www.lfd.uci.edu/~gohlke/pythonlibs/#dpmix
Py-FCM
Download the Windows installer fcm‑0.9.1.win‑amd64‑py2.7.exe from
http://www.lfd.uci.edu/~gohlke/pythonlibs/#py-fcm
Also, download the latest source. This just makes it easier to get the sample file.
https://code.google.com/p/py-fcm/source/browse/
NOTE
I did not install mpi4py, as it is an optional dependency, but here are the links. I also didn’t install the fcm GPU support.
http://www.lfd.uci.edu/~gohlke/pythonlibs/#mpi4py
https://bitbucket.org/mpi4py/mpi4py/downloads
Installation
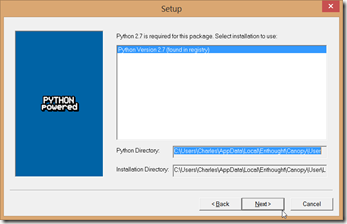
1. Run the Canopy installer. It does not require administrative privileges. I stuck with the defaults.
Clicking “Start using Canopy” will configured the environment. When configured, a window will open, which you can then close. Your desktop will have two icons, one for Canopy, the other for PyLab.
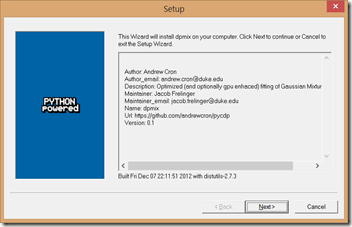
2. Run the DPMix Windows installer. It will find the Python installation and use it.
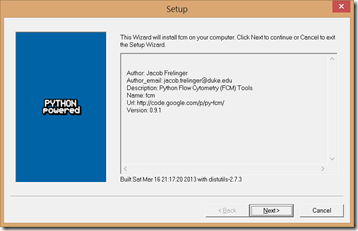
3. Run the py-fcm Windows installer. It likewise will find and use the Python installation.
4. Modify gate.py script. This is critical, as the matplotlib library no longer uses nxutils.
Open this file in a text editor (using your own Username for the directory): C:\Users\Charles\AppData\Local\Enthought\Canopy\User\Lib\site-packages\fcm\core\gate.py
Comment out line 56, and uncomment line 52.
if name is None:
name = self.name
idxs = points_in_poly(self.vert, fcm.view()[:, chan])
# matplotlib has points in poly routine in C
# no faster than our numpy version
#idxs = points_inside_poly(fcm.view()[:, chan], self.vert)
Also comment out the nxutils import on line 2
import numpy
#from matplotlib.nxutils import points_inside_poly
Test
- Unzip the py-fcm source. In the sample_data folder is a file named 3FITC_4PE_004.fcs.
- Open a text editor, and copy the complete path to the file, e.g.
C:\Users\Charles\Documents\FCM Flow Cytometry\Py-FCM\py-fcm-91701cc7a98e\sample_data\3FITC_4PE_004.fcs - Now, replace all back slashes with forward slashes.
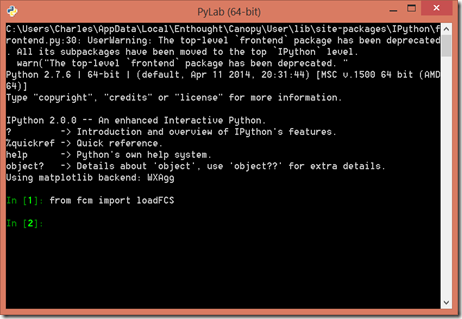
C:/Users/Charles/Documents/FCM Flow Cytometry/Py-FCM/3FITC_4PE_004.fcs - Open PyLab from the icon on the desktop, and run the samples found here (reproduced below).
fcm provides the loadFCS() function to load fcs files:
>>> from fcm import loadFCS
>>> data = loadFCS('../sample_data/3FITC_4PE_004.fcs')
>>> data
3FITC_4PE_004
>>> data.channels
['FSC-H', 'SSC-H', 'FL1-H', 'FL2-H']
>>> data.shape
(94569, 4)
Since the FCMdata object returned by loadFCS() delegates to underlying numpy array, you can pass the FCMdata object to most numpy functions
>>> import numpy as np
>>> np.mean(data)
410.38791252947584
>>> np.mean(data,0)
array([ 538.76464803, 421.57733507, 340.03599488, 341.17367213])
>>> import pylab
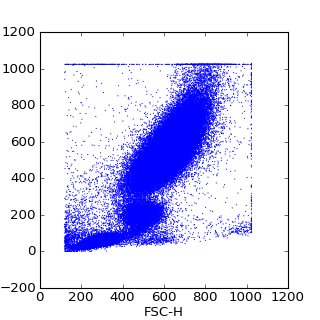
>>> pylab.scatter(data[:,0],data[:,1], s=1, edgecolors='none')
>>> pylab.xlabel(data.channels[0])
>>> pylab.ylabel(data.channels[1])
>>> pylab.show()
References
Info about the nxutiles error. https://groups.google.com/forum/#!topic/py-fcm-dev/HibEajOCTEY
https://pythonhosted.org/fcm/install.html
https://code.google.com/p/py-fcm/
https://www.enthought.com/
http://www.scipy.org/index.html
http://matplotlib.org/
https://github.com/andrewcron/dpmix
http://docs.cython.org/src/quickstart/install.html
https://bitbucket.org/mpi4py/mpi4py/downloads
http://mpi4py.scipy.org/docs/usrman/install.html#requirements